Trna Handles
Background
tRNA can be sequenced with nanopore sequencers, so long as they can be unfolded and electrically attracted to the nanopore. So a mechanism to capture tRNA molecules, unfold them, and initiate threading them into a nanopore is needed.
Technology Description
The invention includes attaching DNA or RNA "handles" to a tRNA molecule. These handles allow manipulation of the tRNA molecule, including unfolding its structure and acting as targets for attaching other molecules to the tRNA.
One example is a double stranded oligonucleotide adapter that can be ligated to a tRNA from a biological sample. Such an adepter can be a Y shaped double stranded DNA-RNA adapter with a 3' RNA overhang complementary to the CCA tail present in tRNA. The adapter can also include a cholesterol tag within its 3' end.
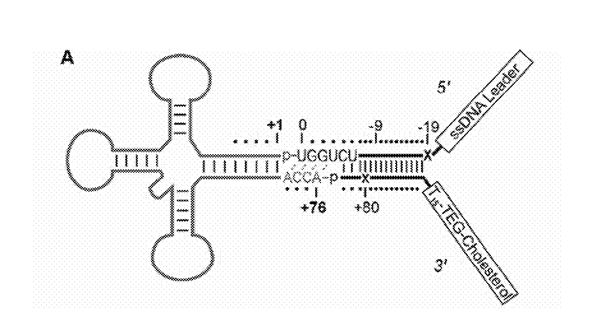
Examples of sequences used in oligonucleotide adaptors include:
GATXGTGAGATCTGATTTTTTTTTTTTTTTZ
GATAGTGAGATCTGATTTTTTTTTTTTTTTZ
GATXGTGAGATCTGATTTTTTTTTTTTTTTZ
X indicates an abasic 1'2' dideoxyribose; Z indicates a triethylene glycol cholesterol
Applications
- Identification of tRNA
- Sequencing of tRNA
- Nanopore sequencing of tRNA
Advantages
- Molecular adaptor that facilitates tRNA seqeuncing by nanopore
- Rapid identification of tRNA species in a biological sample
Intellectual Property Information
| Country | Type | Number | Dated | Case |
| United States Of America | Issued Patent | 10,131,944 | 11/20/2018 | 2014-725 |
| Patent Cooperation Treaty | Published Application | WO 2015/148567 | 10/01/2015 | 2014-725 |
Related Materials
Contact
- University of California, Santa Cruz Industry Alliances & Technology Commercialization
- innovation@ucsc.edu
- tel: View Phone Number.
Inventors
- Bernick, David L.
- Smith, Andrew
Other Information
Keywords
tRNA sequencing, sequencing adapter, tRNA, nanopore sequencing, RNA, RNAseq, long read sequencing
